Hello everyone,
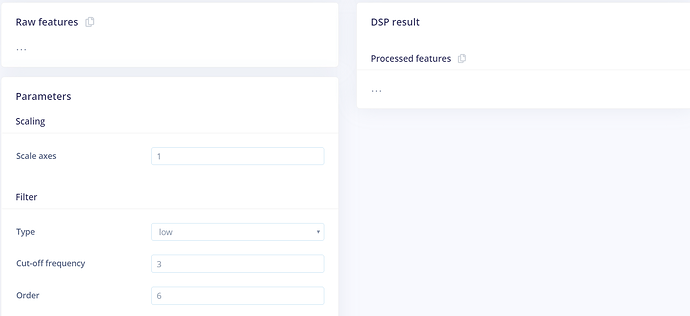
So, I used the ingestion service to get my data and created the default impulse (show in the tutorial video). But, when I go to “Spectral features” Tab, raw features and processed features appear empty.
I’m not sure what is the source of the problem.
Thanks
Hi @doliprane, could you post a screenshot of your impulse configuration? And could you let me know how much data is shown under ‘Data acquisition’?
Hi @janjongboom, so I only used 2 minutes for my example.
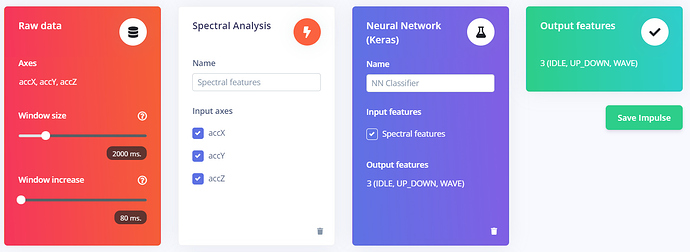
. For the impulse configuration, it’s exactly the one in the tutorial .This is the result in the spectral features tab
@doliprane Could it be that the active sample in spectral features tab is less than 2 seconds? You can select a different sample in the dropdown menu at the top. Also, could you let me know if there are any errors shown in the browser console?
We don’t have a way of logging into your account, so I cannot easily verify myself 
@janjongboom I think that the problem is that I was specifying the wrong interval between two samples. I’m trying to solve this issue right now. I will keep you updated after I test with a new set of samples.
Thank you very much for your assistance 
I tried using a different sample set using different sampling frequencies. I even used 10 ms between every two samples but it’s always the same ouput: no raw features or processed features. only three dots 
@doliprane Is there any error in the browser console? You can also reach out with your credentials to jan@edgeimpulse.com and I’ll take a look directly.
@janjongboom As a matter of fact yes. This error keeps popping multiple times.
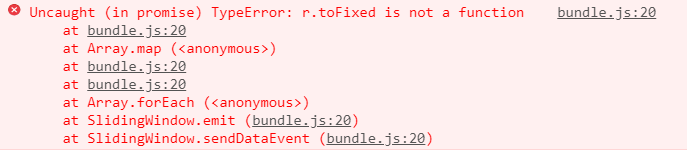
It may be associated to the problem in hand.
Anyway, I sent you my credentials so you can have a closer look at the problem.
I saw it in our logs already, but couldn’t pinpoint what it was  But as said on email, and here for future reference: the values in your ingested messages are strings, not numbers. E.g. see this:
But as said on email, and here for future reference: the values in your ingested messages are strings, not numbers. E.g. see this:
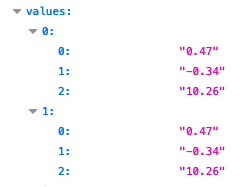
Whereas this should look like:
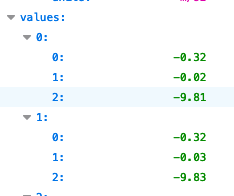
I’ve filed a bug to ensure we throw an error in the ingestion API if we encounter this.
@janjongboom I will make sure to solve the error in my script.
That was really a quick and effcient support and I really appreciate your help 
Thanks a lot